Articles from key opinion leaders
Chronocode, a new concept underlying the control of the biological clock:
The time-keeping mechanism involving rhythmic modifications of the clock proteins

Dr. Yoshitaka Fukada
Professor, Department of Biological Sciences, School of Science,
The University of Tokyo
| 【Biography】 | |
| March 1978 | B.S., Faculty of Science, Kyoto University |
| March 1983 | Ph.D., Graduate School of Science, Kyoto University |
| April 1983 | Assistant Professor, First Department of Biochemistry, Sapporo Medical University School of Medicine |
| September 1986 | Assistant Professor, Department of Biophysics, Faculty of Science, Kyoto University |
| May 1993 | Associate Professor, First Department of Basic Science, College of Arts and Sciences, The University of Tokyo |
| November 1995 – present | Professor, Department of Biological Sciences, School of Science, The University of Tokyo |
| April 1996 – March 1997 | Adjunct Professor, Visiting Researcher, Division of Population Genetics, National Institute of Genetics |
| April 1996 – March 2001 | Principal Investigator, CREST “Understanding the Brain,” JST Strategic Basic Research Programs |
| April 2007 – March 2010 | Program Officer, Biological Sciences Group, Research Center for Science Systems, Japan Society for the Promotion of Science |
| May 2017 – present | Principal Investigator, Grant-in-Aid for Scientific Research of Specially Promoted Research, from MEXT Japan |
1. The biological clock and its research history
“Chronobiology” is the field of biology that studies biological phenomena that change as a function of time, especially phenomena that change in a rhythmic or periodic manner. Phenomena that change unidirectionally over time, such as aging, are not the direct subjects of this field of interests. Most biological rhythms should be controlled by cell-autonomous biological clocks. Among them, the “circadian clock” is the clock mechanism that controls the circadian rhythm with a period of approximately one day (24 hours), which the time required for the Earth to complete one rotation. In most organisms on the earth, many physiological functions have circadian rhythms, and the animal behaviors and physiologies including activity, sleep, body temperature, blood pressure, endocrine secretion, and autonomic function, are governed by the internal circadian clock. Indeed, the influence of the circadian clock is so widespread that almost no physiological function circumvents its control.
The concept of a biological clock was proposed in the 1930s. In support of the concept, Drosophila per mutants with abnormal eclosion rhythms were isolated in 1971. The cloning of the per gene 13 years later revealed the first known clock gene, which led to an explosive era of clock gene discovery. Notably, a significant number of the clock genes were identified by Japanese researchers. An important feature of the circadian clock is that it synchronizes to the light-dark cycle of the external environment. As such, circadian rhythm research has evolved as an extension of two overlapping research fields, chronobiology and photobiology (Figure 1). The common interests of the many excellent researchers in Japan in these two fields created fertile ground for the development of circadian clock research.
Figure 1. Circadian rhythm research evolved as an extension of two overlapping fields, chronobiology and photobiology
No clock in the circadian clock system fails to adapt to the 24-hour light-dark cycle. The circadian clock provides living organisms with the ability to predict the next daily cycle.
2. Transcriptional feedback control of the clock genes and three mysteries
As a clearer picture of clock genes emerged, a feedback mechanism was proposed to explain clock oscillation; the clock gene products negatively regulated their own transcription. At the time, three circadian clock mysteries drew the attention of researchers. The first mystery was how the very long, 24-hour cycle of the oscillation could continue steadily for weeks. This is called “robustness” of the circadian clock. On the other hand, the clock is highly responsive to a variety of environmental signals and could undergo a variable phase shift of the rhythms. The molecular mechanism responsible for this flexibility is the second mystery. The third mystery is a question of how the temperature-sensitivity of the period was suppressed to maintain the roughly 24-hour periodicity of the clock despite ambient temperature changes. This feature is called “temperature compensation” of the period.
3. A new concept on the peripheral clocks
Around 2000, two new concepts emerged that provided a key to solving the three mysteries. One is the concept of peripheral clocks. The body is comprised of countless cellular clocks arranged in a hierarchical coupling system. Because the center of the circadian clock mechanism resides in the hypothalamic suprachiasmatic nucleus, the clock function was originally thought to be a specific cellular function of neurons. However, transcriptional oscillation of the clock genes was seen even in cultured cells, as well as in various tissues and organs in the body. Importantly, the mechanism of the oscillation in the peripheral clock was found to be essentially the same as that of the central clock in the suprachiasmatic nucleus. Thereafter, the clock mechanism was understood as a ubiquitous cellular function present in virtually most of the cells in the body, and the term “peripheral clock” was invented to distinguish satellite clocks from the central one in the suprachiasmatic nucleus. The body is now thought to be comprised of numerous clock cells, and one of the keys to understanding the mystery of clock system stability has been their “coupling.”
4. A new concept of rhythmic phosphorylation that generate 24-hour molecular oscillation
Another new concept is based on the discovery in cyanobacteria that rhythmic phosphorylation of a clock protein, KaiC, is crucial to the clock oscillation. The transcriptional regulation rhythms of the clock genes had been considered as the primary mechanism of the clock oscillation, and all clock oscillation mechanisms proposed in various organisms consisted of two coupling loops as shown in Figure 2. Originally, the transcription-translation feedback loop (on the left, Figure 2) was considered as the primary mechanism of rhythmic oscillation. The phosphorylation cycle of the clock proteins (the loop on the right, Figure 2) was considered a subloop for lengthening circadian periodicity to 24 hours, or a supplementary loop for stabilization of circadian periodicity.
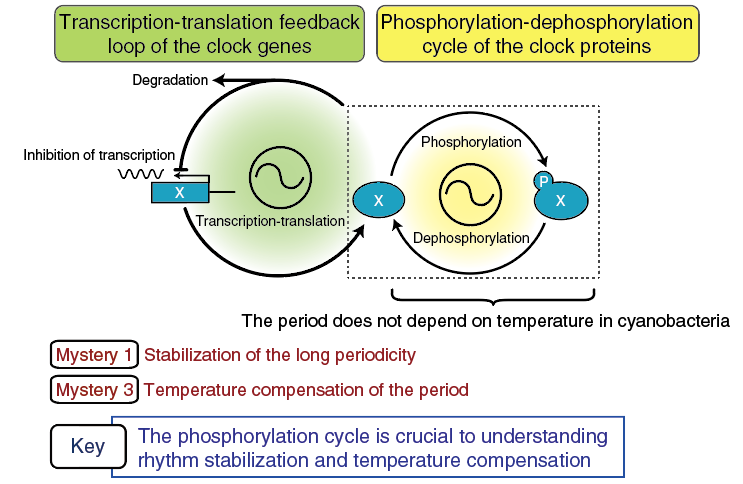
Figure 2. Significance of rhythmic phosphorylation of clock proteins
From cyanobacteria to humans, all clock oscillation mechanisms consist of a transcription feedback loop and phosphorylation cycle.
On the contrary, the basis for the circadian clock oscillation in cyanobacteria was found to be the autophosphorylation-dephosphorylation cycle of KaiC protein alone. Thus, phosphorylation rhythms can be considered the central clock oscillatory mechanism. Although it remains unknown whether such a mechanism is applicable to other clock systems in animals and plants, the point to be emphasized here is that the period of the molecular clock generated by the autophosphorylation rhythms of KaiC is temperature compensated. The phosphorylation cycle may hold the key to understanding the third circadian clock mystery—the mechanism of temperature compensation of the clock period—in animal and plant cells as well. Elucidating the complex regulatory mechanism of the phosphorylation cycle, in addition to the mechanism of rhythm stabilization, is one of the most important subjects for the clock researchers in the future.
We recently found that TGF-beta/activin receptor-like kinase (ALK) [1] and Ca2+/calmodulin-dependent protein kinase (CaMKII) [2] play important roles in the coupling of clock cells. Thus, phosphorylation of clock proteins may also be involved in the hierarchical organization of the clock system.
5. Network of post-translational modifications of the clock proteins: What is the “Chronocode” concept?
As illustrated in Figure 3, nearly all clock proteins are phosphorylated. In many cases, their phosphorylation levels exhibit circadian rhythms. In addition to phosphorylation, many clock proteins undergo various post-translational modifications, such as ubiquitination, acetylation, SUMOylation, and methylation. These modifications play important roles in the regulation of the period and phases of the circadian clock.
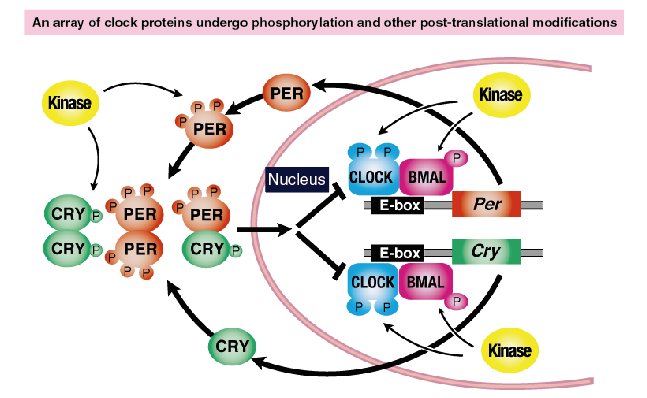
Figure 3. Chronocode: Spatiotemporal regulation of post-translational modifications of the clock proteins
Nearly all clock proteins are regulated by rhythmic phosphorylation by multiple kinases.
We recently discovered two E3 ligases that ubiquitinate the clock protein CRY. They have antagonizing effects, one facilitating the degradation of CRY and the other stabilizing it. Each of these E3 ligases makes the clock oscillate faster or slower, and the loss of both destabilizes the oscillation, leading to arrhythmicity [3]. Ubiquitination of just one clock protein is regulated even by a rather complex regulatory network.
By extrapolation, complex combinations of phosphorylation and other post-translational modifications likely regulate the amount, subcellular localization, and function of an array of clock proteins, helping the clock maintain its periodicity. In other words, the time of day is defined by the modification status of each amino acid in the clock proteins. Based on this idea, I propose the concept of “Chronocode,” which relates to the spatiotemporal regulation of post-translational modifications of the clock proteins. The coined word does not infer a passive description of a time stamp that looks as if encoded by the modification pattern. Instead, these combinations of modifications (“Codes”) that change with time (“Chrono”) are thought to be active participants in maintaining accurate time, or rather define the time-of-the-day. Accordingly, the dynamic nature of this oscillating system could be understood by further characterization of the complex combination of these dynamic modifications of the clock proteins. To give an example, patients with familial advanced sleep-phase syndrome have a significantly advanced sleep phase, and experience wakefulness late at night and sleepiness in the afternoon. This disorder is caused by a point-substitution of a base in the human Per2 DNA leading to a point mutation of an amino acid. This amino acid is a phosphorylation site of PER2 protein, underscoring strongly the significance of spatiotemporal regulation of phosphorylation.
6. Elucidating the chronocode using clock protein antibodies
In the “Chronocode” research, antibody reagents are indispensable tools for high sensitivity detection of an array of clock proteins. Detecting post-translational modifications is often difficult with typical antibodies against proteins. Antibody detection of a band shift due to phosphorylation or ubiquitination could provide the impetus for further experiments. We have generated high-sensitivity monoclonal antibodies specific for CLOCK and BMAL1 [4]. Using these antibodies, we demonstrated that the complex of these transcription factors was phosphorylated and functionally regulated by JNK [5] and CaMKII [2]. With a ChIP analysis using the CLOCK antibody, we identified approximately 8,000 CLOCK binding sites genome-wide where transcription is activated by CLOCK (complexed with BMAL1) in an E-box-dependent manner [6]. By generating CRY antibody and phospho-CRY antibody, we were able to demonstrate that CRY2 was sequentially phosphorylated by DYRK1A and GSK3, leading to degradation [7, 8]. Ordered phosphorylation such as CRY protein is not uncommon in other research fields. We expect that the “Chronocode” concept will spark a Renaissance in the clock research in the near future, once the whole picture of spatiotemporal regulation of post-translational modifications is elucidated by using specific antibodies as an excellent tool.
References
1. Kon et al. (2008) Nature Cell Biol. 10: 1463.
2. Kon et al. (2014) Genes Dev. 28: 1101.
3. Hirano et al. (2013) Cell 152: 1106
4. Yoshitane et al. (2009) Mol. Cell. Biol. 29: 3675.
5. Yoshitane, Honma et al. (2012) EMBO Rep. 13: 455.
6. Yoshitane et al. (2014) Mol. Cell. Biol. 34: 1776.
7. Kurabayashi et al. (2010) Mol. Cell. Biol. 30: 1757.
8. Hirano et al. (2014) Mol. Cell. Biol. 34: 4464.